#include <madness/world/MADworld.h>#include <madness/mra/mra.h>#include <madness/mra/funcplot.h>#include <exception>#include <iterator>#include <list>#include <madness/misc/info.h>#include <madness/chem/AC.h>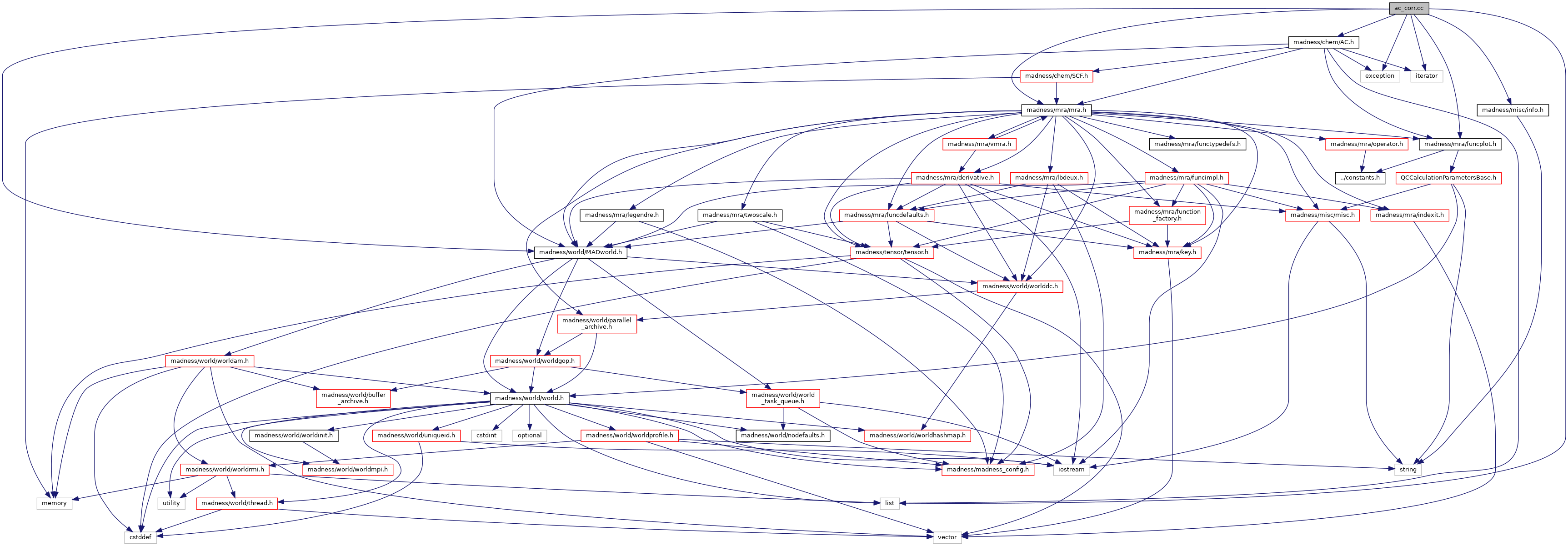
Classes | |
| class | xc_functor< NDIM > |
| Functor for the exchange correlation potential. More... | |
Functions | |
| int | main (int argc, char **argv) |
Function Documentation
◆ main()
| int main | ( | int | argc, |
| char ** | argv | ||
| ) |
interval limits for interpolation region
Coordinates of atoms
Vector for the atom information for all atoms of the molecule
atom information atom 1
put atoms in atom_information vector
Create ACParameters object (necessary to use AC class)
create an object of the AC class to calculate the correction
make xc_potential 1D WARNING: You have to change the code of the density function in the XC functor class to get the right density for your molecule
apply correction
difference between corrected and uncorrected standard potential
References madness::AC< NDIM >::apply(), madness::atom_information< NDIM >::charge, SafeMPI::COMM_WORLD, madness::atom_information< NDIM >::coord, madness::finalize(), madness::initialize(), k, L, param, madness::plot_plane(), madness::print(), madness::atom_information< NDIM >::R1, madness::atom_information< NDIM >::R2, madness::World::rank(), madness::FunctionDefaults< NDIM >::set_cubic_cell(), madness::FunctionDefaults< NDIM >::set_k(), madness::FunctionDefaults< NDIM >::set_thresh(), madness::FunctionDefaults< NDIM >::set_truncate_mode(), madness::slater_radius(), madness::startup(), and thresh.